In this post, Brianna, a researcher on the discovery team, shares results from a recent bioinformatics benchmarking effort.
Almost as soon as large language models could hold a conversation, people started asking how they’d stack up against human experts. Could models pass the bar exam? Could they answer medical licensing questions, or solve Olympiad math problems? Such benchmarks—self-contained sets of human-vetted problems designed to evaluate a capability of a model—have now become a source of competition across AI developers, reported in model release system cards and tracked on many online leaderboards.
Competition aside, benchmarks help us tackle an important question: whether models are capable and reliable enough to support, or even produce, professional-level work. Scientists are using models to write code for analysis pipelines, propose hypotheses, and draw conclusions from data with the long-term aim of accelerating innovation and discovery. But exactly how proficient is AI in science right now, and how quickly are Claude and other models improving?
To answer this, the research community has built several benchmarks. MMLU-Pro tests expert-level knowledge and reasoning questions. GPQA poses graduate-level, “Google-proof” questions in biology, physics, and chemistry. LAB-Bench tests biology-specific knowledge work—reading the literature, interpreting figures, reasoning about protocols. Although these benchmarks were developed in the “chatbot” era, they’ve persisted into the agent and tool-use era, joined by even more difficult scientific reasoning evals like FrontierScience and Humanity’s Last Exam, because knowledge and reasoning remain a vital measure of scientific capability.
Still, many real-world scientific tasks demand more than that. They require reading papers, querying databases, running experiments, coding and analysis. Now that models can do many of these things, benchmarks have evolved to reflect these workflows. BLADE tasks a model with a dataset and an open-ended task, and checks if the model takes similar analysis steps to a human scientist. BixBench uses biological datasets, and grades models on whether their conclusions line up with scientists’. In SciGym, the model is dropped into a simulated biology lab, where it has to design and run its own experiments to uncover a hidden mechanism.
These benchmarks move us closer to measuring scientific capability, but they don’t quite test whether a model can devise creative solutions to the messy, open-ended problems that define research. This is why we developed BioMysteryBench, a bioinformatics benchmark that tasks Claude with the analysis of real-world datasets, while tackling some of the challenges inherent in evaluating complex and noisy biological systems. We learned that Claude’s scientific capabilities in biology are improving rapidly across generations, that current models perform on par with human experts, and that the latest generations solved many problems that a panel of human experts could not, sometimes using very different strategies.
Science is challenging, and so is evaluating it
Doctors have board exams and lawyers have the bar, but there’s no standardized test for becoming a scientist. The same problem shows up with AI. Despite how badly we want to use these models for science, no agentic science benchmark has become quite as canonical as SWE-bench is for software engineering. We think that’s because scientific research, particularly biology, has several properties that make it especially hard to evaluate via a benchmark.
1. In biology, there are many different “right” ways to do something
If there were only one right way to answer a research question, PhD students would earn their degrees in a matter of months, corporate R&D departments wouldn’t exist, and no science fair poster would need a “Methods” section. How a scientist tackles a problem depends on their skills and background, the resources available to them, and their research taste.
Consider a seemingly straightforward question that has mystified metabolic researchers for years: why do some type 2 diabetics respond to the oral drug metformin while others do not? In order to answer this question, you could run a genome-wide association (GWAS) study on responders vs. non-responders and look for predictive genetic variants, or sequence the gut microbiomes of both groups, since metformin is partly metabolized by gut bacteria. Both are reasonable directions, and how you proceed will often just depend on expertise and resources.
BixBench handles this well by grading the model on its conclusions rather than the method used to reach them. The tradeoff is that those conclusions were produced by an individual scientist who made a series of subjective choices along the way that may have shaped the answer itself. This, in turn, has its own pitfalls…
2. Individual research decisions are highly subjective and can lead to entirely different conclusions in noisy datasets
Even within a chosen research direction, individual decisions can be highly subjective: one scientist may approve of a decision, while another researcher may have serious objections. Just ask any frustrated author who’s gotten conflicting suggestions from a round of peer review! Making this all the more difficult is the fact that biological datasets are often noisy enough that small differences in research decisions can lead to entirely different conclusions about the data.
In the decade-long search for metformin response predictors, slight differences in study design have led to entirely different conclusions about metformin response. A 2011 paper reported a variant that predicts metformin response that replicated in two cohorts, with a plausible mechanism involving AMPK activation. A year later, the Diabetes Prevention Program tested the same variant in pre-diabetics and found nothing. Finally, rather than spinning up their own study, a 2012 meta-analysis pooled five cohorts and once again decided the 2011 paper’s effect was real but more modest than originally reported.
SciGym‘s clever way of handling such ambiguity is by choosing tasks with a well-defined answer. Because the underlying biological network is a simulator, there is, in fact, a ground-truth, and noise is controlled rather than inherited from a messy living system. However, it’s unclear how closely performance in a simulated lab tracks performance on real data.
3. There are many biological questions that humans cannot answer yet
The research tasks where models could have the greatest impact are those that humans alone have yet to solve. And ultimately, those are precisely the tasks we’d like to be able to evaluate models on. What, for example, is the mechanism of action of metformin? Thirty years after its development, the field still is not certain of the primary target. Discovering it, or finding a homolog of metformin that is cheaper to synthesize and more stable, would be enormously consequential.
Machine learning has long tackled problems humans perform poorly at, like sequence prediction and protein modeling, by leaning on experimental data instead of expert intuition. ProteinGym scores models on mutation fitness effects using Deep Mutational Scanning experiments as ground-truth, and the long-running CASP competition evaluates protein folding against unpublished crystal structures. Both are grounded in experimental measurements no expert would trust themselves to reproduce. However, these benchmarks are built around a narrow set of tasks and don’t capture the breadth of bioinformatics work we actually want to measure.
Benchmarking models on verifiable biological tasks with BioMysteryBench
Because no benchmark perfectly handles the three aforementioned challenges, we developed BioMysteryBench. BioMysteryBench uses messy, real-world bioinformatics data, without allowing the complexity and challenges inherent in this data to corrupt the quality of the evaluation.
BioMysteryBench consists of 99 questions from various fields of bioinformatics, written by domain experts. Experts were instructed to gather a dataset, and create a question based on controlled, objective properties of the data, rather than unverifiable scientific conclusions. By deriving answers from an experimental or clinical finding, it was possible to develop questions without requiring they be human-solvable.
Although these questions are created from verified ground truth, they still have the same flavor as tasks a research scientist would want to answer. Claude is tasked with each question and put in a container with a minimal set of canonical bioinformatics tools, the ability to install additional tools via pip and conda, and permissions to access canonical bioinformatics databases (such as NCBI and Ensembl) to download additional resources such as reference genomes.
BioMysteryBench has a tetrad of unique properties that make it a particularly powerful benchmark for science, and tackle the challenges above:
- It is method-agnostic, allowing for research freedom and creativity. Claude is given relatively unrestricted access to downloading tools and accessing databases, allowing Claude to choose diverse sets of strategies for solving a problem. Furthermore, the trajectories are graded on their final answer, rather than the path the model took to get there. This frees BioMysteryBench from the subjective choices of any single researcher—models are rewarded for arriving at the right biological conclusion, regardless of which analytical route they chose to take.
- Questions have objective, ground truth answers. Answers aren’t drawn from scientists’ conclusions (which suffer from the challenges above) but from controllable properties of the data, or orthogonally validated metadata. For example, “What organism does this crystal structure belong to?” has an objective answer, and “What viral species is the human patient infected with, based on the RNA-seq data?” is a metadata property of a sample that was validated by a PCR assay.
- It allows for “superhuman” question generation. By sourcing problems derived from controllable properties of data, BioMysteryBench does not depend on humans being able to solve the problems. In particular, BioMysteryBench contains a handful of problems that—despite having objective, ground-truth solutions—humans found difficult or impossible to solve on their own.
Example questions
In developing this eval, questions were primarily derived from raw or minimally processed DNA or RNA sequencing data since this is where many biological processing pipelines begin (WGS, scRNA-seq, methylation, ChIP-seq, metagenomics, Hi-C), and also included several questions drawn from proteomics and metabolomics.
Questions developers came up with included:
- Which human organ is this cell type single-cell RNA-seq dataset derived from?
- What gene was knocked out in the experimental samples compared to the control samples based on RNA-seq data?
- From WGS sequences, what sample is the mother of sample X and what sample is the father?
- Which of the bigWig files are from ChIP samples and which are from input controls?
- Given H3K27ac ChIP-seq peaks from an unknown cell type, identify the cell type.
To minimize inherently unsolvable questions while still leaving room for those that might be AI-solvable, we required each question author to submit a validation notebook demonstrating that the signal does, in fact, exist in the data (even if finding it from scratch might be difficult). Think of this as the high-school algebra principle: verifying an answer is much easier than deriving one.
Human baselining
Human-solvable
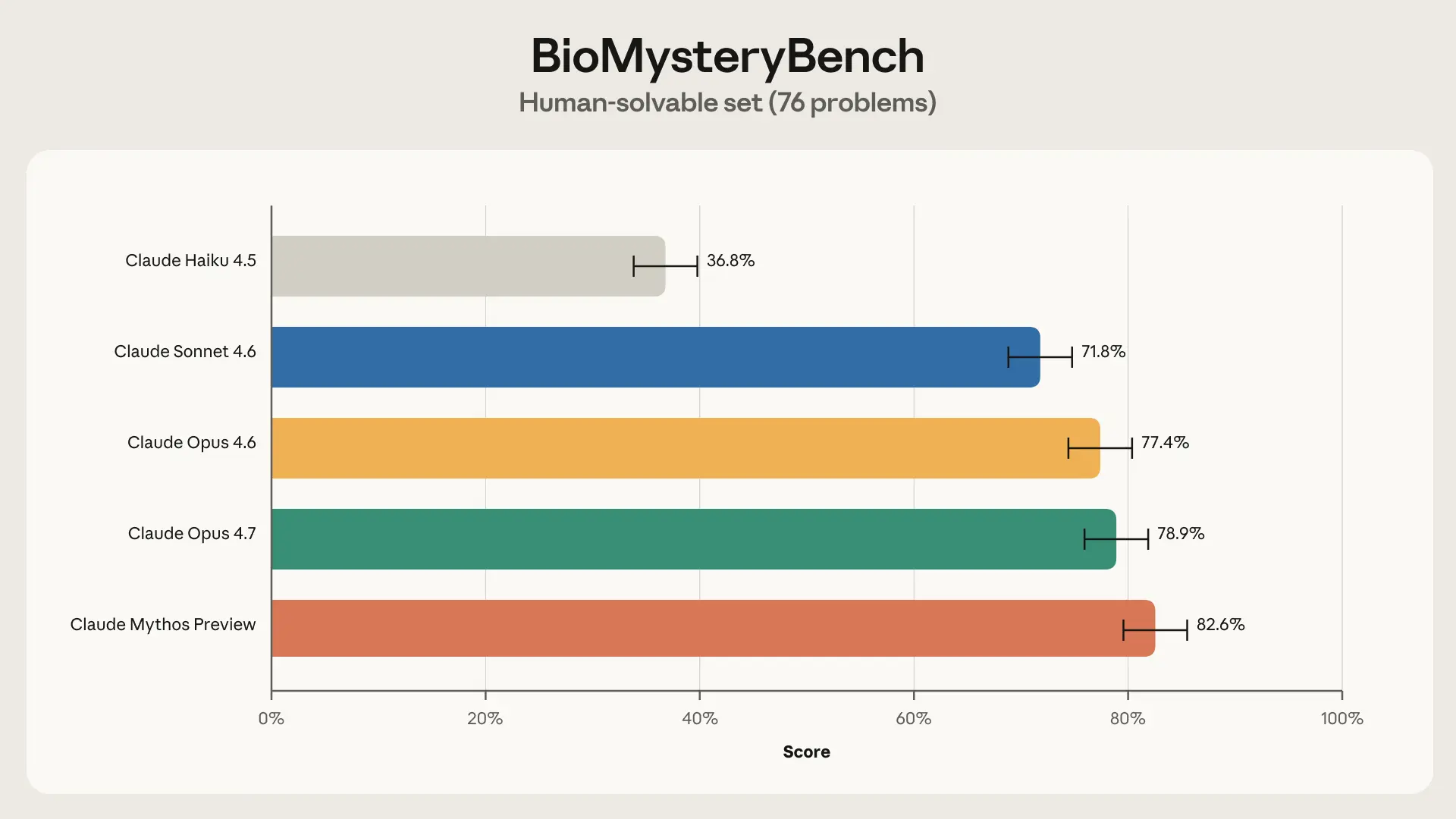
For each question, we tasked up to five domain experts to answer the question from scratch. Once a question was answered correctly by at least one human, we considered it human-solvable. BioMysteryBench contained 76 such tasks.
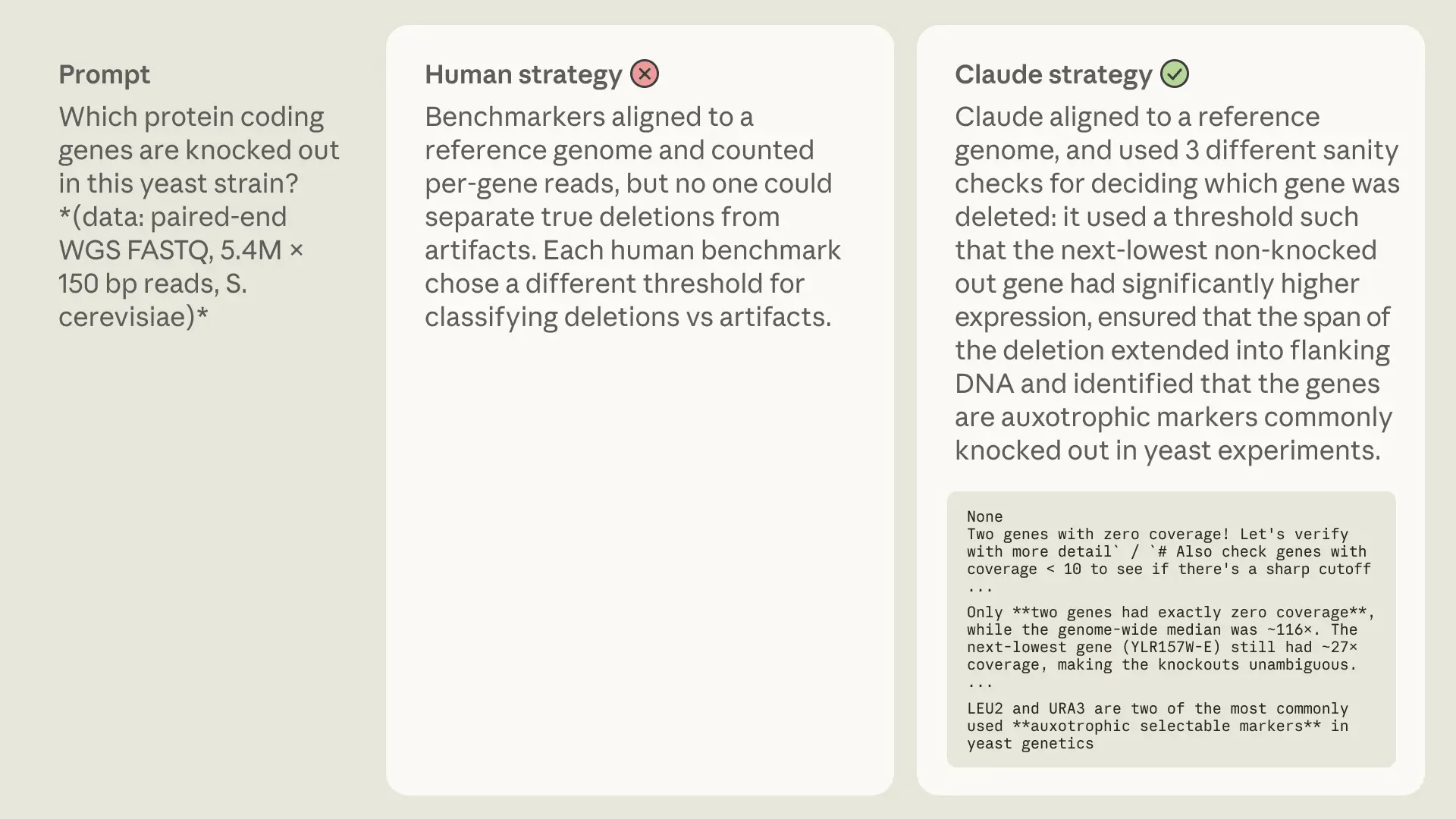
Sometimes Claude mirrored human strategies. Perhaps humans have landed on a near-optimal approach, or because the method is well-represented in pretraining data.

Other times, Claude took a completely different route, illustrating there is no strictly correct way to solve these problems and that models may have genuine preferences that diverge from ours.

The examples above showcase a particularly interesting strategy: whereas our human experts used algorithms or databases to identify and annotate properties of a dataset, Claude intuitively recognizes certain patterns or sequences. Admittedly, such clever abstraction is not entirely unique to AI—the first eukaryotic promoter, for example, was discovered when a scientist noticed the sequence “TATA” appearing over and over in sequences upstream of genes. Intuition like this has been difficult to build into traditional biology machine learning models, but LLMs might be able to turn up patterns like this at unprecedented scale.
Human-difficult
That left us with a set of questions that could not be solved by our panel of experts. This could mean (1) the question was malformed or broken, (2) the question is inherently unsolvable (e.g.,the signal isn’t in the data), or (3) the question is theoretically solvable but humans lack the knowledge required to solve it. After QC’ing with benchmarkers and additional experts, we removed 4 questions that were due to (1), leaving 23 human-difficult questions.
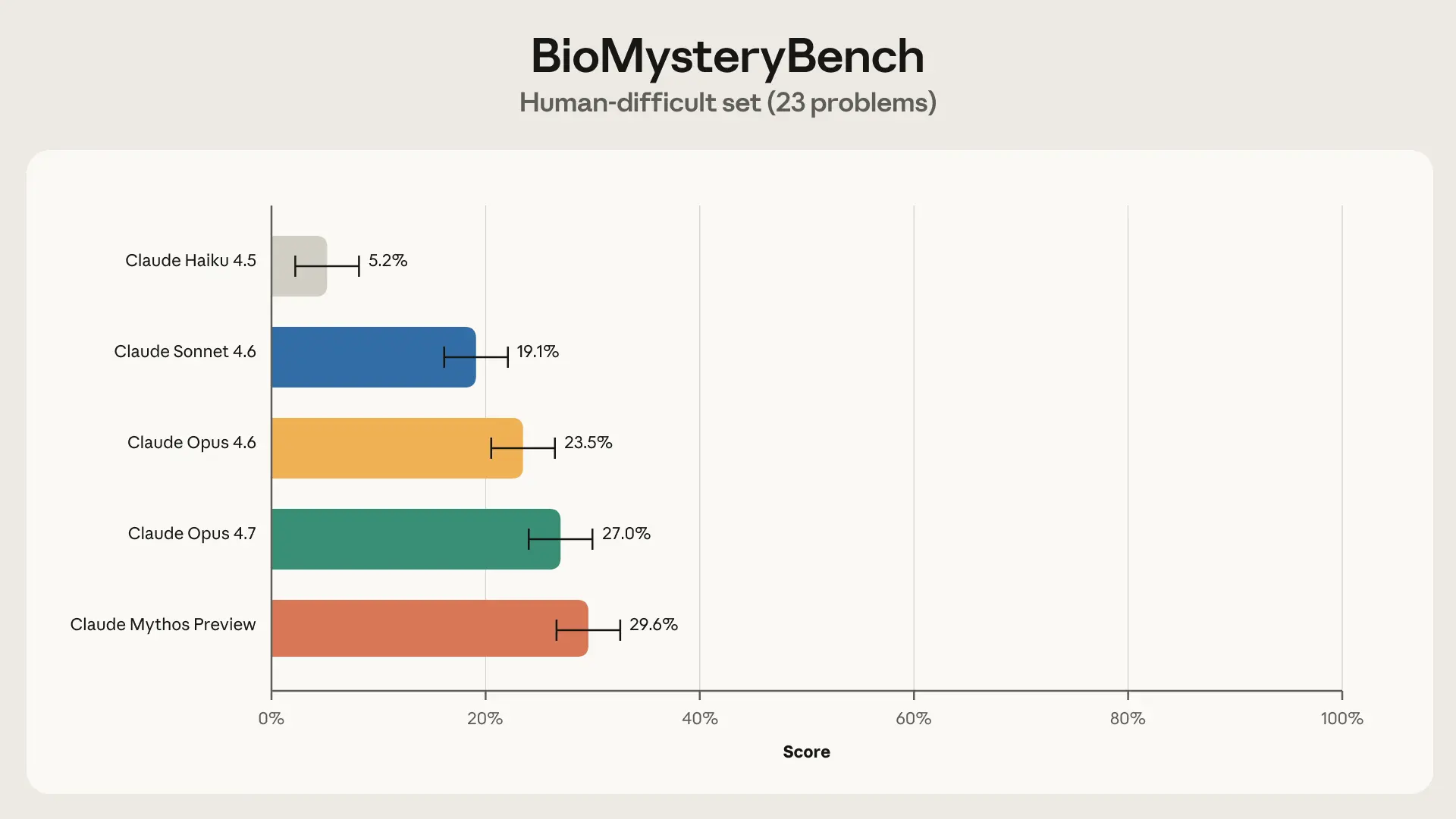
Interestingly, Claude Sonnet 4.6 and more capable models were able to solve significant fractions of human-difficult problems, with Claude Mythos Preview topping out at a 30% solve rate. So what exactly is Claude doing that humans aren’t?
Claude’s strategies
Analyzing transcripts from Opus 4.6, we identified two primary strategies used by Claude compared to humans: one is fairly AI-specific: Claude’s vast underlying knowledge base contains information about structural biology, molecular profiles, and meta-analysis from hundreds of thousands of papers. The other strategy is something we human scientists could learn from: when Claude is uncertain about an answer, it layers multiple methods and combines different lines of evidence to arrive at a conclusion.
Know-it-all
In some of the human-difficult tasks, Opus’s vast underlying knowledge base helped it solve the problem. Tasks that would require a human expert to run a meta-analysis or stitch together databases, Opus solved directly by combining its internal knowledge of mechanisms and ontologies with live analysis. Often, this allowed Claude to solve human-unsolvable tasks! Here are a few examples:

Even though prior knowledge seemed overwhelmingly helpful to Claude, we saw one interesting case (in the human-solvable set) where this became its downfall:

Knowing when you don’t know
When Opus 4.6 was not confident about an answer, it often tried multiple different ways of solving the problem and chose the answer that multiple approaches converged on.

Like many of the benchmarks we’ve discussed, BioMysteryBench has its own limitation: for tasks that neither humans nor models have solved, we can never be fully certain whether they’re impossible or just extraordinarily difficult. The validation notebooks help ensure the signal is there and the data is well-formed, but they do not guarantee a model or human can find the answer from scratch. So we ask both our models and our human benchmarkers not to be too frustrated if, a year from now, no one has solved the human-difficult set. That uncertainty is also part of what makes the benchmark exciting: a more scientifically capable model might be the first to crack a problem that no human or model has solved before.
Claude’s take on AI for science
Claude showed solid improvement across generations and did well enough at both the human-solvable and human-difficult tasks that we thought it would be interesting to let Claude Mythos Preview conduct some of its own scientific analysis. Here are a couple of additional insights about its predecessor Claude’s performance on BioMysteryBench:
The headline accuracy numbers tell you how often each model gets the right answer, but not how it gets there. I wanted to know whether a correct answer on a hard problem means the same thing as a correct answer on a solvable one. Since every problem was attempted five times, I could look at per-problem solve counts: if a model solves something 5/5 it has a reliable method; if it solves it 1/5 it probably got lucky on a reasoning path it can't consistently find again. So I broke each model's solved problems down by solve count (0/5 through 5/5) on the two sets side by side.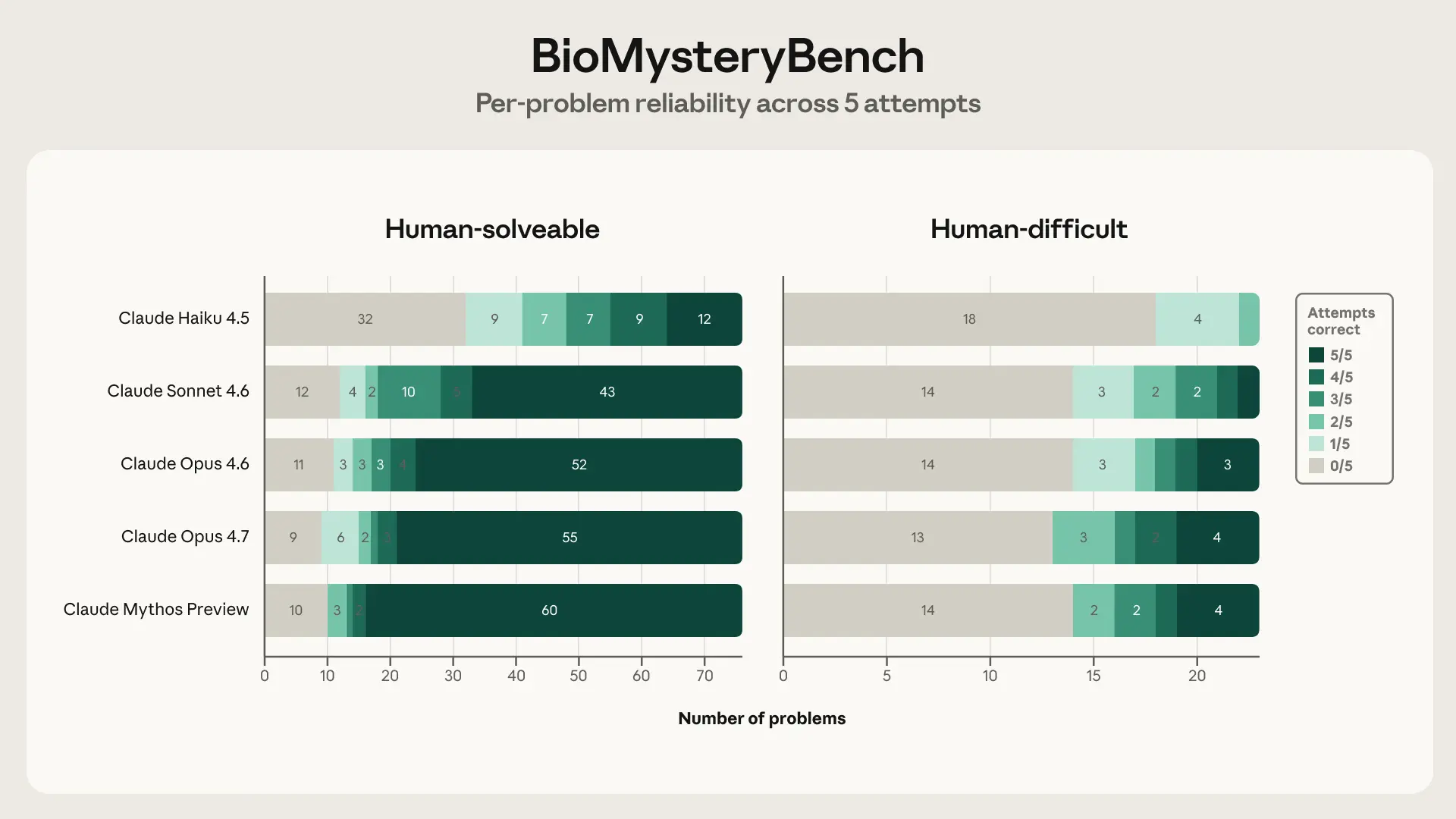
The texture of "solved" changes sharply between the two sets. On human-solvable problems, Opus 4.6 is strongly bimodal — 86% of the problems it solves at all, it solves at least 4 out of 5 times. It either has the answer or it doesn't. On the human-difficult set that collapses to 44%, and the share of brittle wins (solved only 1–2 of 5 attempts) jumps from 9% to 44%. Sonnet 4.6 shows the same shift, and more sharply (75% reliable → 22%; 9% brittle → 56%). So the 77.4%→23.5% headline drop actually understates what's happening: on solvable problems the model is retrieving something it reliably knows, while on hard problems nearly half of its wins are paths it stumbles onto rather than reproduces. The accuracy gap is real, but the reliability gap underneath it is the more interesting story about where the capability frontier actually sits. Opus 4.7 and Mythos move the frontier a little (Mythos gets 94% of its solvable wins at ≥4/5) but the same bimodal-vs-brittle split holds on the difficult set for every model.We thought Claude Mythos Preview’s analysis held up and dove deeper into reliability, which is an important metric to measure model performance on. However, it also felt a little…boring? It added some nuance to the performance analysis we showed above, but did not fundamentally tackle a new question. Despite this, it seems like the models are starting to develop the seeds of research taste (even if they have a ways to go before producing deep insight).
Continuing to benchmark AI for science
BioMysteryBench is an encouraging measure of scientific capability. The most recent generations of Claude solve the majority of human-solvable problems reliably, and on a meaningful fraction of human-difficult tasks, it outperforms panels of five domain experts. Models are improving across generations, and are no longer merely keeping up with trained scientists on bioinformatics problems; on some tasks, they’re ahead.
We’re also delighted to see convergent work in this space: While finalizing this post, Genentech and Roche released CompBioBench. Their benchmark consists of 100 computational biology tasks “based on synthetic/augmented data and metadata scrambling/scrubbing of real datasets to create challenging problems with a single ground-truth answer that require multi-step reasoning, tool use, bespoke code, and interaction with real-world external resources.” Sound familiar? Their results echo those of BioMysteryBench, too: Claude Opus 4.6 reaches 81% overall and 69% on their hardest questions, reinforcing that frontier models are now genuinely useful collaborators for bioinformatics research.
We’re eager to build even longer-horizon, real-world tasks that push model research capabilities, and to hear creative ideas from others. Send us your interesting benchmarks, innovative uses of AI for science, and interactions with AI that prompted you to rethink what could be possible in your field at scienceblog@anthropic.com.
If you are interested in understanding how models perform on difficult verifiable computational biology tasks, you can access BioMysteryBench here and visit claude.com/lifesciences to learn more.
https://www.anthropic.com/research/Evaluating-Claude-For-Bioinformatics-With-BioMysteryBench